The genome of the novel SARS-CoV-2 codes for an ORF1a/ ORF1ab (open reading frame) polyprotein containing sixteen non-structural proteins (NSP) and four structural proteins. The genome also has multiple ORFs coding for accessory proteins through a frame shift. These accessory proteins are not necessary for viral replication but might play a key role in pathogenesis of SARS-CoV-2. One such protein is the accessory protein 7a, which is predicted to contribute to Covid-19 by inducing the apoptotic processes in human host cells1.
Structure
SARS-CoV-2 is a very young virus and the structure and function of the accessory protein 7a has not yet been solved. However, 7a of SARS-CoV-2 shows 85% sequence identity and 95.2% sequence similarity with another protein in SARS-CoV2. It is therefore conceivable that both accessory proteins have a similar structure and function. The sequence analysis of SARS-CoV predicts that ORF7a codes for a type I transmembrane protein with 122 amino acids, including a signal peptide at the N‑terminus and a retrieval signal at the C-terminus3. The N-terminal ectodomain of ORF7a consists of seven β-strands compactly arranged in an immuno-globulin-like β-sandwich fold (Fig 1). These seven β-strands are ordered in two β-sheets containing four β-strands (A; G; F; C) in the first sheet and three (B; E; D) in the second one (see Fig 1: left)4.
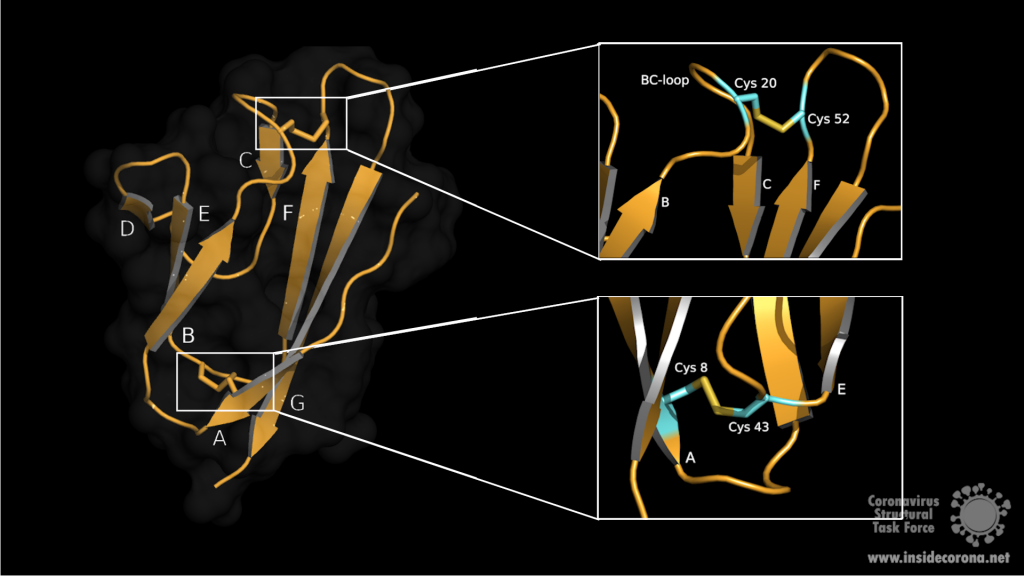
Both sheets are amphipathic, with the hydrophobic side facing inwards closely packed against each other. The top of the ectodomain is defined by the BC, DE and FG loops and the bottom by the AB, CD and EF loops. The β-sandwich structure is stabilized by two disulphide bonds linking the sheets at opposite edges. At the bottom of the structure, a disulphide bridge connects Cys8 on strand A with Cys43 at the end of strand E. At the top, Cys20 of the BC loop is linked to Cys52 at the end of strand F (see Fig 1: right). Additionaly, on top of the BED sheet, the DE loop protrudes from the structure and forms a groove together with β-strands C and D. In the centre is Glu18 which contributes to the negatively charged bottom of the mainly hydrophobic groove. This grove may be a potential site for ligand interaction due to its central negative electrostatic potential4.
Function
In cell culture, the polypeptide 7a of SARS-CoV seems to have diverse biological functions5. It is possible that 7a plays a key role in cell cycle control. In HEK 293 cells, an overexpression of 7a led to inhibition of cell growth and induction of the G0/G1 phase cell cycle arrest. This arrest may favour coronavirus replication and exacerbate virus-induced pathogenicity. 7a is also predicted to induce apoptosis in human kidney epithelial cells by interaction with a protein called B-cell lymphoma-extra large (Bcl-XL). Bcl-XL belongs to a group of pro-survival proteins, the B-cell lymphoma-2 (Bcl-2)- family, which prevent apoptosis in epithelial cells. The Interaction between 7a and the C-terminal transmembrane domain of Bcl-XL may interfere with this pro-survival function, leading to apoptosis via the caspase-dependant pathway6,7. In addition to this, SARS 7a interacts with a Ap4A-hydrolase involved in cell proliferation, DNA-replication, apoptosis and RNA-processing. This interaction leads to downregulation of its hydrolase-activity and an increased production of AP4A (diadenosine tetraphosphate) which may also induce apoptosis5. Such a host cell specific modulation of apoptosis could enable the virus to evade the immune response or to spread to other target organs.
Another predicted function of ORF7a is the inhibition of the bone marrow matrix antigen 2 (BST-2) that might restrict virus release by physically tethering the budding enveloped virion to the plasma membrane. ORF7a antagonizes this function by binding of the extracellular domain of BST-2 preventing its glycosylation. Thus, an inhibitor preventing ORF7a-BST-2 interaction can be speculated as potential drug target8.
Taken together, ORF7a is a virulence factor that contributes in different ways to the pathogenicity of SARS-CoV-2. Therefore, targeted drug development against ORF7a could be a critical factor to reduce viral spread or attenuate severe disease progression.
PDB Structures Available
6W37: X-ray structure of the SARS-CoV-2 ORF7a encoded accessory protein.
1xak: SARS-CoV ORF7a accessory protein, a unique type I transmembrane protein of unknown function. Has a short cytoplasmic tail and a transmembrane domain. Consists of one chain (chain A), that forms a compact seven-stranded beta sandwich.
1y04: SARS Coronavirus ORF 7a coded X4 protein, also known as 7a, U122 or X4. Type-I transmembrane protein with immunoglobulin like beta-sandwich fold. Potential functions of X4 in virus replication and pathogenesis are discussed.
References
- 1.Michel CJ, Mayer C, Poch O, Thompson JD. Characterization of accessory genes in coronavirus genomes. Virol J. Published online August 27, 2020. doi:10.1186/s12985-020-01402-1
- 2.Francis K. Y. The Proteins of Severe Acute Respiratory Syndrome Coronavirus‑2 (SARS CoV‑2 or n‑COV19), the Cause of COVID‑19. The Protein Journal (2020). 2020;(39):198-216. doi:10.1007/s10930-020-09901-4
- 3.Fielding BC, Tan Y-J, Shuo S, et al. Characterization of a Unique Group-Specific Protein (U122) of the Severe Acute Respiratory Syndrome Coronavirus. JVI. Published online July 15, 2004:7311-7318. doi:10.1128/jvi.78.14.7311-7318.2004
- 4.Hänel K, Stangler T, Stoldt M, Willbold D. Solution structure of the X4 protein coded by the SARS related coronavirus reveals an immunoglobulin like fold and suggests a binding activity to integrin I domains. J Biomed Sci. Published online November 23, 2005:281-293. doi:10.1007/s11373-005-9043-9
- 5.Vasilenko N, Moshynskyy I, Zakhartchouk A. SARS coronavirus protein 7a interacts with human Ap4A-hydrolase. Virology Journal. Published online 2010:31. doi:10.1186/1743-422x-7-31
- 6.Tan Y-J, Fielding BC, Goh P-Y, et al. Overexpression of 7a, a Protein Specifically Encoded by the Severe Acute Respiratory Syndrome Coronavirus, Induces Apoptosis via a Caspase-Dependent Pathway. JVI. Published online December 15, 2004:14043-14047. doi:10.1128/jvi.78.24.14043-14047.2004
- 7.Tan Y-X, Tan THP, Lee MJ-R, et al. Induction of Apoptosis by the Severe Acute Respiratory Syndrome Coronavirus 7a Protein Is Dependent on Its Interaction with the Bcl-XL Protein. JVI. Published online April 11, 2007:6346-6355. doi:10.1128/jvi.00090-07
- 8.Taylor JK, Coleman CM, Postel S, et al. Severe Acute Respiratory Syndrome Coronavirus ORF7a Inhibits Bone Marrow Stromal Antigen 2 Virion Tethering through a Novel Mechanism of Glycosylation Interference. García-Sastre A, ed. J Virol. Published online September 16, 2015:11820-11833. doi:10.1128/jvi.02274-15

[…] of SARS-CoV-2 VUI-202012/01 has 14 amino acid changes and three deletions affecting the genes for ORF1ab, spike and ORF8. One of these mutations (N501Y) occurs in the receptor binding domain and could […]